-
PDF
- Split View
-
Views
-
Cite
Cite
Nomura Tatsuji, Stephen Potkay, Establishment and Preservation of Reference Inbred Strains of Rats For General Purpose Use: Report on U.S. – Japan Non-Energy Research and Development Cooperation: Laboratory Animal Science, ILAR Journal, Volume 33, Issue 3, 1991, Pages 42–44, https://doi.org/10.1093/ilar.33.3.42
Close - Share Icon Share
Introduction
Inbred strains of rats are becoming increasingly important in biomedical research. Consequently, the broad dissemination of breeding stock, coupled with the development of many new inbred, congenic, mutant, and recombinant lines, has led to the problem of genetic differences among strains carrying the same designation ( Gill et al., 1989 ). Indeed, Bender et al. (1984 ), Natori et al. (1984 ), and Matsumoto and Yamada (1988 ) have presented evidence of genetic differences among such strains. Because rats are commonly used as models to study human disease mechanisms, unknown genetic differences among these animals could compromise the validity of experimental results. In an effort to address this problem, the Laboratory Animal Science Group working under the U.S.--Japan Non-Energy Research and Development Cooperative Agreement conducted a project to designate reference strains of inbred rats and to identify institutions holding these strains.
Rationale for Establishing Reference Strains
There may be several reasons for the appearance of strains with the same designation but different genotypes. These include residual heterozygosity resulting from the counteracting effects of natural selection for reproductive fitness, independent derivation from the same outbred stock, separation before complete genetic fixation, mutation, and unplanned outcrossing
Selection of Reference Rat Strains
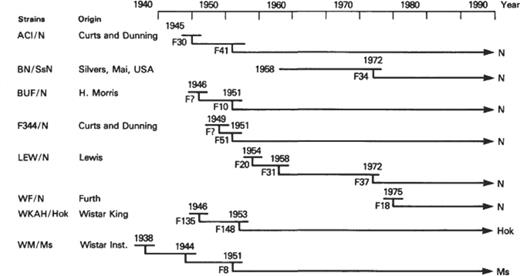
Eight established rat strains were selected to serve as reference strains. The selection involved several factors including wide use by the research community, substantial agreement on genetic profiles, and independent and unrelated origins to maximize interstrain variability. Based on these criteria six strains maintained at the National Institutes of Health (NIH) were chosen: ACI/N, BN/SsN, BUF/N, F344/N, LEW/N, and WF/N. Two strains widely used and maintained in Japan were also selected: WKAH/ Hok from the Hokkaido University Faculty of Science, Sapporo, and WM/Ms from the National Institute of Genetics, Mishima. The origins of these strains are shown in Figure 1 . Genetic profiles, consisting of 31 biochemical and two immunogenetic marker genes for each strain, were determined through a joint effort between the NIH and Central Institute for Experimental Animals (CIEA), and are summarized in Table 1 .
Allele Distribution of Reference Inbred Strains of Rats
| . | Biochemical loci . | hnmunogenetic loci . | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Acon . | Acp . | Ahd . | Ahd . | Akp . | Alp . | Amy . | Cs . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Fh . | Gc . | Gdc . | Gst . | Hao . | Hbb . | Lap . | Mup . | Pep . | Pg . | Rtp . | Rtp . | Svp . | RT1 . | RT2 . |
| Strains . | -1 . | -2 . | -2 . | -c . | -1 . | -1 . | -1 . | -1 . | -1 . | -2 . | -3 . | -4 . | -6 . | -7 . | -8 . | -9 . | -10 . | -14 . | -1 . | . | -1 . | -1 . | -1 . | . | -1 . | -1 . | -3 . | -1 . | -1 . | -2 . | -1 . | . | . |
| ACI/N | b | a | c | a | b | -- | b | a | b | a | a | b | b | b | b | a | a | a | b | a | a | b | a | b | b | b | a | a | b | a | a | a | b |
| BN/SsN | a | -- | b | b | a | b | b | a | a | c | d | b | -- | b | a | c | b | a | a | a | b | a | b | a | a | a | b | a | a | a | b | n | a |
| BUF/N | b | a | c | a | a | b | a | a | b | a | a | b | a | b | b | a | a | a | b | b | a | b | a | b | a | b | a | b | b | b | b | b | a |
| F344/N | b | a | c | a | a | b | a | -- | a | a | a | b | a | b | b | a | a | b | b | a | a | b | a | a | a | b | b | b | b | a | a | l | a |
| LEW/N | b | a | c | b | a | b | a | a | a | d | d | b | a | b | a | c | b | b | a | a | a | b | a | b | b | b | a | a | b | b | b | l | a |
| WF/N | a | a | a | b | a | -- | a | a | a | c | c | b | b | b | a | c | b | b | b | a | a | b | a | a | a | b | a | a | b | b | b | u | b |
| WKAH/Hok | b | a | c | -- | a | a | a | b | a | a | a | b | -- | b | b | a | a | b | b | -- | -- | b | a | b | -- | -- | -- | -- | -- | -- | a | k | a |
| WM/Ms | b | a | a | a | a | b | a | b | a | d | d | b | a | b | a | c | b | b | b | a | -- | -- | a | b | -- | b | -- | b | -- | -- | a | u | b |
| . | Biochemical loci . | hnmunogenetic loci . | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Acon . | Acp . | Ahd . | Ahd . | Akp . | Alp . | Amy . | Cs . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Fh . | Gc . | Gdc . | Gst . | Hao . | Hbb . | Lap . | Mup . | Pep . | Pg . | Rtp . | Rtp . | Svp . | RT1 . | RT2 . |
| Strains . | -1 . | -2 . | -2 . | -c . | -1 . | -1 . | -1 . | -1 . | -1 . | -2 . | -3 . | -4 . | -6 . | -7 . | -8 . | -9 . | -10 . | -14 . | -1 . | . | -1 . | -1 . | -1 . | . | -1 . | -1 . | -3 . | -1 . | -1 . | -2 . | -1 . | . | . |
| ACI/N | b | a | c | a | b | -- | b | a | b | a | a | b | b | b | b | a | a | a | b | a | a | b | a | b | b | b | a | a | b | a | a | a | b |
| BN/SsN | a | -- | b | b | a | b | b | a | a | c | d | b | -- | b | a | c | b | a | a | a | b | a | b | a | a | a | b | a | a | a | b | n | a |
| BUF/N | b | a | c | a | a | b | a | a | b | a | a | b | a | b | b | a | a | a | b | b | a | b | a | b | a | b | a | b | b | b | b | b | a |
| F344/N | b | a | c | a | a | b | a | -- | a | a | a | b | a | b | b | a | a | b | b | a | a | b | a | a | a | b | b | b | b | a | a | l | a |
| LEW/N | b | a | c | b | a | b | a | a | a | d | d | b | a | b | a | c | b | b | a | a | a | b | a | b | b | b | a | a | b | b | b | l | a |
| WF/N | a | a | a | b | a | -- | a | a | a | c | c | b | b | b | a | c | b | b | b | a | a | b | a | a | a | b | a | a | b | b | b | u | b |
| WKAH/Hok | b | a | c | -- | a | a | a | b | a | a | a | b | -- | b | b | a | a | b | b | -- | -- | b | a | b | -- | -- | -- | -- | -- | -- | a | k | a |
| WM/Ms | b | a | a | a | a | b | a | b | a | d | d | b | a | b | a | c | b | b | b | a | -- | -- | a | b | -- | b | -- | b | -- | -- | a | u | b |
Alleles at biochemical loci were decided by electrophoresis; those at immunogenetic loci were decided by a hemagglutination test using antisear.
--: Not tested.
Allele Distribution of Reference Inbred Strains of Rats
| . | Biochemical loci . | hnmunogenetic loci . | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Acon . | Acp . | Ahd . | Ahd . | Akp . | Alp . | Amy . | Cs . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Fh . | Gc . | Gdc . | Gst . | Hao . | Hbb . | Lap . | Mup . | Pep . | Pg . | Rtp . | Rtp . | Svp . | RT1 . | RT2 . |
| Strains . | -1 . | -2 . | -2 . | -c . | -1 . | -1 . | -1 . | -1 . | -1 . | -2 . | -3 . | -4 . | -6 . | -7 . | -8 . | -9 . | -10 . | -14 . | -1 . | . | -1 . | -1 . | -1 . | . | -1 . | -1 . | -3 . | -1 . | -1 . | -2 . | -1 . | . | . |
| ACI/N | b | a | c | a | b | -- | b | a | b | a | a | b | b | b | b | a | a | a | b | a | a | b | a | b | b | b | a | a | b | a | a | a | b |
| BN/SsN | a | -- | b | b | a | b | b | a | a | c | d | b | -- | b | a | c | b | a | a | a | b | a | b | a | a | a | b | a | a | a | b | n | a |
| BUF/N | b | a | c | a | a | b | a | a | b | a | a | b | a | b | b | a | a | a | b | b | a | b | a | b | a | b | a | b | b | b | b | b | a |
| F344/N | b | a | c | a | a | b | a | -- | a | a | a | b | a | b | b | a | a | b | b | a | a | b | a | a | a | b | b | b | b | a | a | l | a |
| LEW/N | b | a | c | b | a | b | a | a | a | d | d | b | a | b | a | c | b | b | a | a | a | b | a | b | b | b | a | a | b | b | b | l | a |
| WF/N | a | a | a | b | a | -- | a | a | a | c | c | b | b | b | a | c | b | b | b | a | a | b | a | a | a | b | a | a | b | b | b | u | b |
| WKAH/Hok | b | a | c | -- | a | a | a | b | a | a | a | b | -- | b | b | a | a | b | b | -- | -- | b | a | b | -- | -- | -- | -- | -- | -- | a | k | a |
| WM/Ms | b | a | a | a | a | b | a | b | a | d | d | b | a | b | a | c | b | b | b | a | -- | -- | a | b | -- | b | -- | b | -- | -- | a | u | b |
| . | Biochemical loci . | hnmunogenetic loci . | |||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | Acon . | Acp . | Ahd . | Ahd . | Akp . | Alp . | Amy . | Cs . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Es . | Fh . | Gc . | Gdc . | Gst . | Hao . | Hbb . | Lap . | Mup . | Pep . | Pg . | Rtp . | Rtp . | Svp . | RT1 . | RT2 . |
| Strains . | -1 . | -2 . | -2 . | -c . | -1 . | -1 . | -1 . | -1 . | -1 . | -2 . | -3 . | -4 . | -6 . | -7 . | -8 . | -9 . | -10 . | -14 . | -1 . | . | -1 . | -1 . | -1 . | . | -1 . | -1 . | -3 . | -1 . | -1 . | -2 . | -1 . | . | . |
| ACI/N | b | a | c | a | b | -- | b | a | b | a | a | b | b | b | b | a | a | a | b | a | a | b | a | b | b | b | a | a | b | a | a | a | b |
| BN/SsN | a | -- | b | b | a | b | b | a | a | c | d | b | -- | b | a | c | b | a | a | a | b | a | b | a | a | a | b | a | a | a | b | n | a |
| BUF/N | b | a | c | a | a | b | a | a | b | a | a | b | a | b | b | a | a | a | b | b | a | b | a | b | a | b | a | b | b | b | b | b | a |
| F344/N | b | a | c | a | a | b | a | -- | a | a | a | b | a | b | b | a | a | b | b | a | a | b | a | a | a | b | b | b | b | a | a | l | a |
| LEW/N | b | a | c | b | a | b | a | a | a | d | d | b | a | b | a | c | b | b | a | a | a | b | a | b | b | b | a | a | b | b | b | l | a |
| WF/N | a | a | a | b | a | -- | a | a | a | c | c | b | b | b | a | c | b | b | b | a | a | b | a | a | a | b | a | a | b | b | b | u | b |
| WKAH/Hok | b | a | c | -- | a | a | a | b | a | a | a | b | -- | b | b | a | a | b | b | -- | -- | b | a | b | -- | -- | -- | -- | -- | -- | a | k | a |
| WM/Ms | b | a | a | a | a | b | a | b | a | d | d | b | a | b | a | c | b | b | b | a | -- | -- | a | b | -- | b | -- | b | -- | -- | a | u | b |
Alleles at biochemical loci were decided by electrophoresis; those at immunogenetic loci were decided by a hemagglutination test using antisear.
--: Not tested.
Maintenance, Production, and Preservation of Reference Strains
The principal objective in the maintenance and production of these strains is to minimize the probability of genetic changes over many generations. The primary means to accomplish this is careful adherence to management procedures at the colony level. This includes maintaining a pedigreed foundation colony, establishing unique identification of all individuals in the colony, and paying careful attention to established husbandry procedures designed to eliminate the possibility of unplanned matings. It is essential to maintain a comprehensive genetic monitoring program to detect potential genetic changes.
Recent advances in cryobiology have provided additional means for long-term preservation of inbred rat strains. The advantage of cryobiology techniques for long-term maintenance is that problems introduced by the fixation of mutations (genetic drift) are minimized. Cryopreservation programs are underway at NIH and CIEA. Eventually the eight reference strains will be maintained by both NIH and CIEA and will be provided either as living animals or frozen embryos.
Availability of Reference Strains
Small numbers of animals representing the reference strains are available for establishing breeding colonies. Requests should be addressed to Chief, Scientific Services Branch, Veterinary Resources Program, National Center for Research Resources, National Institutes of Health, Building 14A, Room 102, Bethesda, MD 20892 USA; or, Director, Central Institute for Experimental Animals, 1430 Nogawa Miyamae Kawasaki Kanagawa 216, JAPAN
Conclusions
The eight inbred rat strains that have been selected to serve as reference strains are the result of an initiative by the Laboratory Animal Science Group working under the U.S.--Japan Non-Energy Research and Development Cooperative Agreement. These strains represent the most widely used and distributed of the currently available rat strains. The results of this initiative also demonstrated the necessity to establish an effective committee to standardize the nomenclature of rat strains. Efforts are underway to establish such a committee.
References